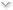The Green Cloud is the invention of Dr. Paul Brenner at the ND Center for Research Computing. It is a containerized data center located at the South Bend city greenhouse, stocked with used servers kindly donated by Ebay, Inc. The first batch of machines was installed in December, and will eventually reach about 400 cores once everything is turned on.

What makes the data center unique is that is has no air conditioning. Instead, the data center takes in ambient air, and then exhausts it into the greenhouse. This benefits Notre Dame, since we no longer pay the cost of cooling, but it also benefits the greenhouse, which has significant heating costs during the winter months. (We used to call this idea grid heating.)
Of course, this means the capacity of the system may change with the weather. During the winter, the system can run at full blast and deliver as much heat as possible to the greenhouse. During the spring and fall, the heat may not be needed, and can be vented outdoors. During the hottest part of the summer, we may need to shut some machines down to get the temperature under control. However, recent studies by big data center operators suggest that machine temperature could be safely increased to 80 or 90 degrees, so there may be a fair amount of headroom available. We will see.
For a normal data center that runs web servers and databases, shutting down machines is not really an option. However, the Green Cloud provides fungible computing power for large computations in science and engineering at Notre Dame. If structured correctly, these workloads can adapt to 10 or 100 or 1000 cores. So, turning machines on and off will affect performance, but not correctness.
A good example of a flexible workload is genome assembly. Two of our students, Christopher Moretti and Michael Olson presented initial results on a Scalable Genome Assembler at the MTAGS Workshop held at Supercomputing 2009. Their assembler uses our Work Queue framework to manage a variable workforce, pushing out sequence fragments to whatever machines are available. The system has scaled up to about 1000 cores, spread across the Notre Dame campus, the Green Cloud, Purdue University, and the University of Wisconsin.
We are currently working on a journal paper and an open source release of the assembler, so stay tuned for details.